DeepSeek has made a lot of noise lately. Their R1 model, released in January 2025, outperformed competitors like OpenAI’s O1 at launch. But what truly set it apart was its highly efficient infrastructure—dramatically reducing costs while maintaining top-tier performance.
Now, they're coming for data engineers. DeepSeek released a bunch of small repositories as independent code modules. Thomas Wolf, Co-founder and Chief of Product at HuggingFace shared some of his highlights, but we're going to focus on one particularly important project went that unmentioned—smallpond, a distributed compute framework built on DuckDB. DeepSeek is pushing DuckDB beyond its single-node roots with smallpond, a new, simple approach to distributed computing.
First, having DeepSeek, a hot AI company, using DuckDB is a significant statement, and we'll understand why. Second, we'll dive into the repository itself, exploring their smart approach to enabling DuckDB as a distributed system, along with its limitations and open questions.
I assume you're familiar with DuckDB. I've created tons of content around it. But just in case, here's a high-level recap.
For transparency, at the time of writing this blog, I’m a data engineer and DevRel at MotherDuck. MotherDuck provides a cloud-based version of DuckDB with enhanced features. Its approach differs from what we’ll discuss here, and while I’ll do my best to remain objective, just a heads-up! 🙂
DuckDB Reminder
DuckDB is an in-process analytical database, meaning it runs within your application without requiring a separate server. You can install it easily in multiple programming languages by adding a library—think of it as the SQLite of analytics, but built for high-performance querying on large datasets.
It's built in C++ and contains all the integrations you might need for your data pipelines (AWS S3/Google Cloud Storage, Parquet, Iceberg, spatial data, etc.), and it's damn fast. Besides working with common file formats, it has its own efficient storage format—a single ACID-compliant file containing all tables and metadata, with strong compression.
In Python, getting started is as simple as:
pip install duckdbThen, load and query a Parquet file in just a few lines:
import duckdb
conn = duckdb.connect()
conn.sql("SELECT * FROM '/path/to/file.parquet'")It also supports reading and writing to Pandas and Polars DataFrames with zero copy, thanks to Arrow.
import duckdb
import pandas
# Create a Pandas dataframe
my_df = pandas.DataFrame.from_dict({'a': [42]})
# query the Pandas DataFrame "my_df"
# Note: duckdb.sql connects to the default in-memory database connection
results = duckdb.sql("SELECT * FROM my_df").df()DuckDB is coming into AI companies?
We talk a lot about LLM frameworks, models, and agents, but we often forget that the first step in ANY AI project comes down to data.
Whether it's for training, RAG, or other applications, it all comes down to feeding systems with good, clean data. But how do we even accomplish that step? Through data engineering. Data engineering is a crucial step in AI workflows but is less discussed because it's less "sexy" and less "new."
Regarding DuckDB, we've already seen other AI companies like HuggingFace using it behind the scenes to quickly serve and explore their datasets library through their dataset viewer.
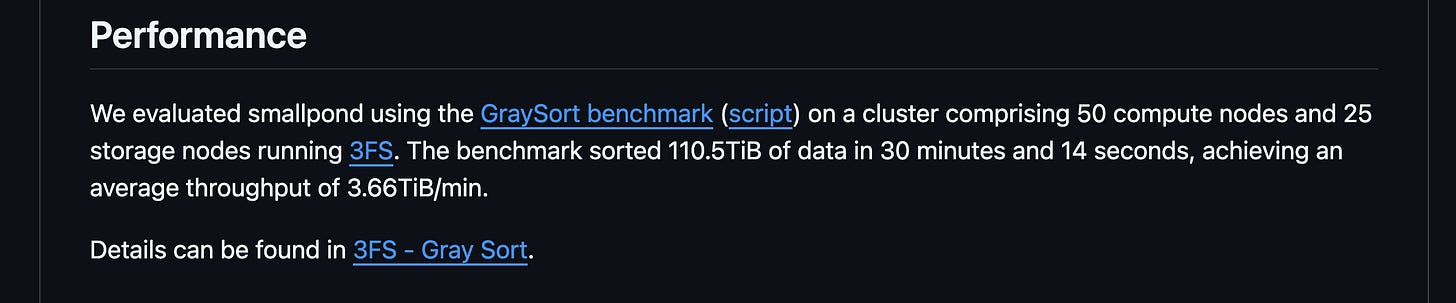
Now, DeepSeek is introducing smallpond, a lightweight open-source framework, leveraging DuckDB to process terabyte-scale datasets in a distributed manner. Their benchmark states: _“Sorted 110.5TiB of data in 30 minutes and 14 seconds, achieving an average throughput of 3.66TiB/min.”
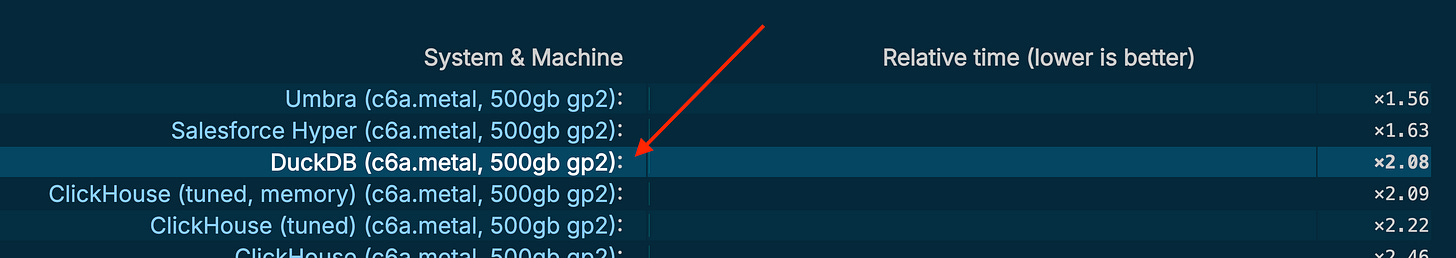
While we've seen DuckDB crushing 500GB on a single node easily (clickbench), this enters another realm of data size.
But wait, isn't DuckDB single-node focused? What's the catch here?
Let's dive in.
smallpond's internals
DAG-Based Execution Model
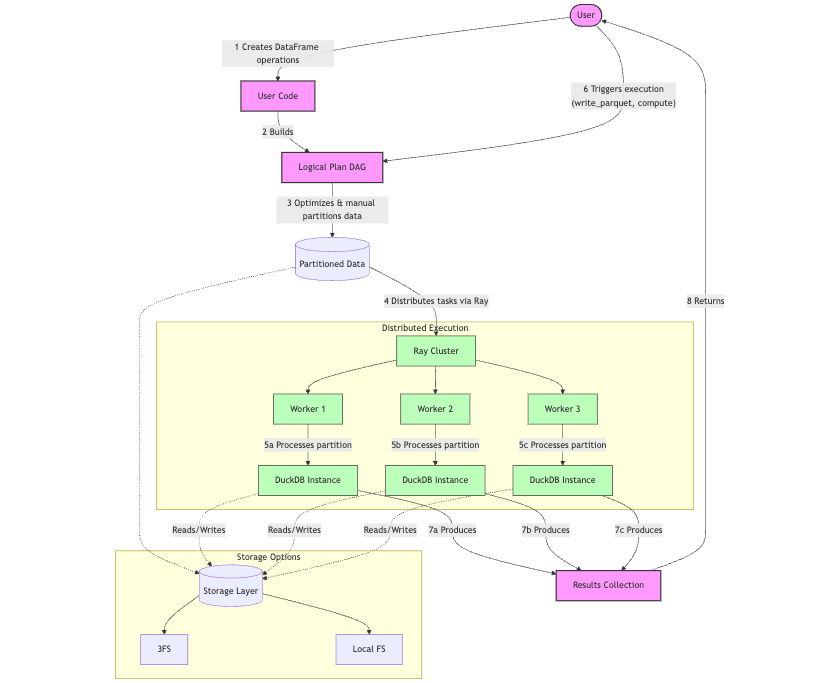
smallpond follows a lazy evaluation approach when performing operations on DataFrames (map(), filter(), partial_sql(), etc.), meaning it doesn’t execute them immediately. Instead, it constructs a logical plan represented as a directed acyclic graph (DAG), where each operation corresponds to a node in the graph (e.g., SqlEngineNode, HashPartitionNode, DataSourceNode).
Execution is only triggered when an action is called, such as:
write_parquet()– Write data to diskto_pandas()– Convert to a pandas DataFramecompute()– Explicitly request computationcount()– Count rowstake()– Retrieve rows
This approach optimizes performance by deferring computation until necessary, reducing redundant operations and improving efficiency.
When execution is triggered, the logical plan is converted into an execution plan. The execution plan consists of tasks (e.g., SqlEngineTask, HashPartitionTask) that correspond to the nodes in the logical plan. These tasks are the actual units of work that will be distributed and executed through Ray.
Ray Core and Distribution Mechanism
The important thing to understand is that the distribution mechanism in smallpond operates at the Python level with help from Ray, specifically Ray Core, through partitions.
A given operation is distributed based on manual partitioning provided by the user. Smallpond supports multiple partitioning strategies:
Hash partitioning (by column values)
Even partitioning (by files or rows)
Random shuffle partitioning
For each partition, a separate DuckDB instance is created within a Ray task. Each task processes its assigned partition independently using SQL queries through DuckDB.
Given this architecture, you might notice that the framework is tightly integrated with Ray, which comes with a trade-off: it prioritizes scaling out (adding more nodes with standard hardware) over scaling up (improving the performance of a single node).
Therefore, you would need to have a Ray cluster. Multiple options exist, but you would have more options today to manage your own cluster through AWS/GCP compute or a Kubernetes cluster. Only Anyscale, the company founded and led by the creators of Ray, offers a fully managed Ray service. Even then, you have the overhead of monitoring a cluster.
The great thing here is that the developer experience is nice because you get a local single node when working and only scale when you need to. But the question is: do you actually need to scale out and add the cluster overhead given that the largest machine on AWS today provides 24TB of memory?
Storage Options
Ray Core is just for compute - where does the storage live?
While smallpond supports local filesystems for development and smaller workloads, the benchmark on 100TB mentioned is actually using the custom DeepSeek 3FS framework: Fire-Flyer File System is a high-performance distributed file system designed to address the challenges of AI training and inference workloads.
To put it simply, compared to AWS S3, 3FS is built for speed, not just storage. While S3 is a reliable and scalable object store, it comes with higher latency and eventual consistency, making it less ideal for AI training workloads that require fast, real-time data access. 3FS, on the other hand, is a high-performance distributed file system that leverages SSDs and RDMA networks to deliver low-latency, high-throughput storage. It supports random access to training data, efficient checkpointing, and strong consistency, eliminating the need for extra caching layers or workarounds. For AI-heavy workloads that demand rapid iteration and distributed compute, 3FS offers a more optimized, AI-native storage layer—trading off some cost and operational complexity for raw speed and performance.
Because this is a specific framework from DeepSeek, you would have to deploy your own the 3FS cluster if you want to reach the same performance. There's no fully managed option there... or maybe this is an idea for a spinoff startup from DeepSeek? 😉
One interesting experiment would be to test performance at the same scale using AWS S3. However, this implementation is currently missing in smallpond. This approach would be much more practical for an average company needing 100TB of processing capability.
Key differences from other frameworks like Spark/Daft
Unlike systems like Spark or Daft that can distribute work at the query execution level (breaking down individual operations like joins or aggregations), smallpond operates at a higher level. It distributes entire partitions to workers, and each worker processes its entire partition using DuckDB.
This makes the architecture simpler but potentially less optimized for complex queries that would benefit from operation-level distribution.
Summary of smallpond's Architecture
Let's recap the features of smallpond :
Lazy evaluation with DAG-based execution – Operations are deferred until explicitly triggered.
Flexible partitioning strategies – Supports hash, column-based, and row-based partitioning.
Ray-powered distribution – Each task runs in its own DuckDB instance for parallel execution.
Multiple storage layer options – Benchmarks have primarily been conducted using 3FS.
Cluster management trade-off – Requires maintaining a compute cluster, though fully managed services like Anyscale can mitigate this.
Potential 3FS overhead – Self-managing a 3FS cluster introduce significant additional complexity.
Other ways of distributed compute with DuckDB
Another approach to distributed computing with DuckDB is through serverless functions like AWS Lambda. Here, the logic is often even simpler than partitioning, typically processing by file. Or you could decide to process per partition with some wrapper, but you won't be able to go much further than file-by-file processing.
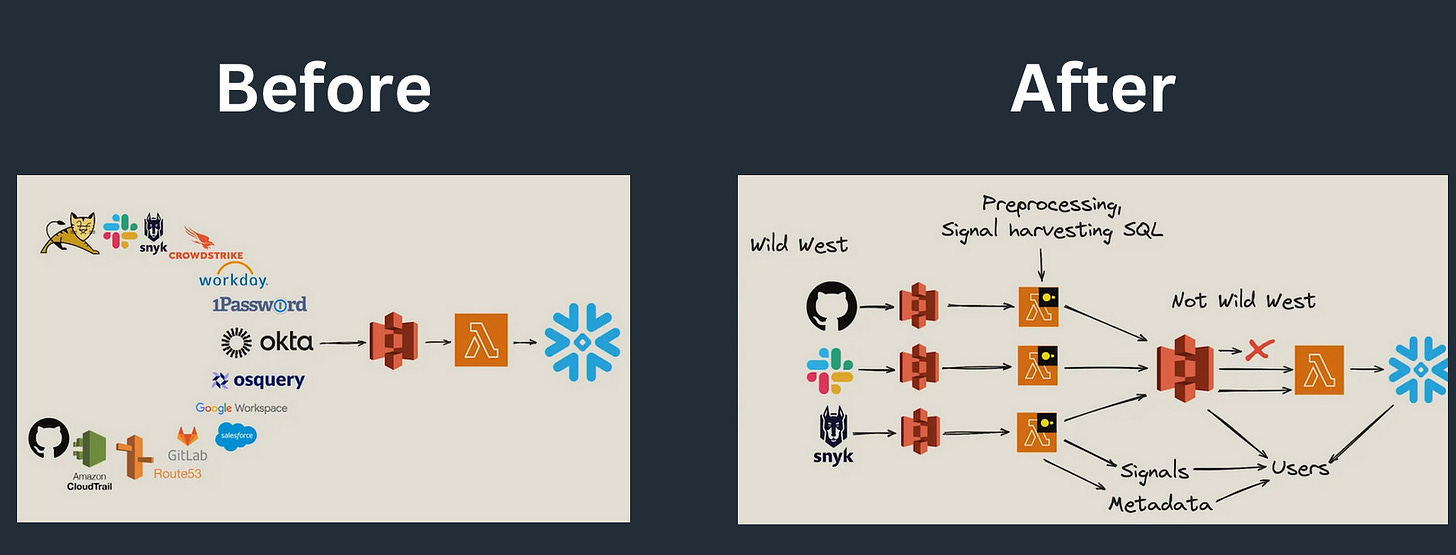
Okta implemented this approach, and you can read more on blog: Okta's Multi-Engine Data Stack
Finally, MotherDuck, is working on dual execution, balancing between local and remote compute for optimize resources usage.
Scaling DuckDB
All in all, it's exciting to see that DuckDB is being used in AI-heavy workloads and that people are getting creative on how to split the compute when needed.
smallpond, while being restricted to a specific tech stack for distributing compute, aims to be simple, which aligns with the philosophy of DuckDB 👏
It's also a good reminder that there are multiple ways to scale DuckDB. Scaling up is always the simpler approach, but with smallpond and other examples mentioned here, we have plenty of options.
This approach makes sense nowadays rather than having to rely on complex and heavy distributed frameworks by default "just in case." These not only hurt your cloud costs when starting with small/medium data but also has a tax on developer experience (still love you, Apache Spark ❤️).
While we have powerful single-node solutions that would be enough for most use cases, especially if you're in the 94% of use cases under 10TB according to Redshift, we now have even more options to make the Duck fly.